Note
Go to the end to download the full example code.
Comparing methods for approximating the hypervolume#
The following examples compare various ways of approximating the hypervolume of a nondominated set.
Comparing HypE and Rphi-FWE+#
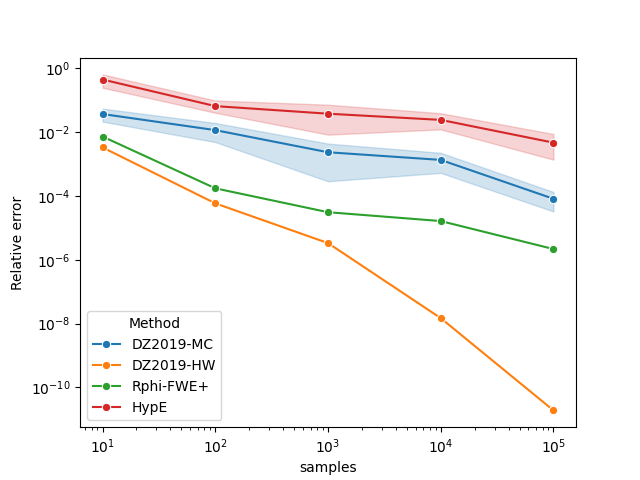
This example shows how to approximate the hypervolume metric of the
CPFs.txt dataset using both moocore.whv_hype() (HypE), and
moocore.hv_approx() for several values of the number of
samples between \(10^1\) and \(10^5\). We repeat each calculation 5
times to account for stochasticity.
import time
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import moocore
First calculate the exact hypervolume.
ref = 2.1
x = moocore.get_dataset("CPFs.txt.xz")[:, :-1]
x = moocore.filter_dominated(x)
x = moocore.normalise(x, to_range=[1, 2])
true_hv = moocore.hypervolume(x, ref=ref)
Next, we approximate the hypervolume using \(\{10^1, 10^2, \ldots, 10^5\}\) random samples to show the higher samples reduce the approximation error. Since the approximation is stochastic, we perform 5 repetitions of each computation.
nreps = 5
nsamples_exp = 5
rng1 = np.random.default_rng(42)
rng2 = np.random.default_rng(42)
results = []
for i in range(1, nsamples_exp + 1):
for r in range(nreps):
res = moocore.whv_hype(x, ref=ref, ideal=0, nsamples=10**i, seed=rng1)
results.append(dict(Method="HypE", rep=r, samples=i, value=res))
res = moocore.hv_approx(
x, ref=ref, nsamples=10**i, seed=rng2, method="DZ2019-MC"
)
results.append(dict(Method="DZ2019-MC", rep=r, samples=i, value=res))
res = moocore.hv_approx(x, ref=ref, nsamples=10**i, method="DZ2019-HW")
results.append(dict(Method="DZ2019-HW", rep=0, samples=i, value=res))
res = moocore.hv_approx(x, ref=ref, nsamples=10**i, method="Rphi-FWE+")
results.append(dict(Method="Rphi-FWE+", rep=0, samples=i, value=res))
width = len("Mean DZ2019-MC")
text = "True HV"
print(f"{text:>{width}} : {true_hv:.5f}")
df = pd.DataFrame(results)
df["Method"] = (
df["Method"]
.astype("category")
.cat.reorder_categories(["DZ2019-MC", "DZ2019-HW", "Rphi-FWE+", "HypE"])
)
subdf = (
df[df["samples"] == 4]
.groupby("Method", observed=True)["value"]
.agg(["mean", "min", "max"])
)
for what, mean, min, max in subdf.itertuples():
text = "Mean " + what
print(f"{text:>{width}} : {mean:.5f} [{min:.5f}, {max:.5f}]")
True HV : 1.05704
Mean DZ2019-MC : 1.05766 [1.05579, 1.06019]
Mean DZ2019-HW : 1.05704 [1.05704, 1.05704]
Mean Rphi-FWE+ : 1.05703 [1.05703, 1.05703]
Mean HypE : 1.07701 [1.04252, 1.11705]
Next, we plot the results.
df["samples"] = 10 ** df["samples"]
df["value"] = np.abs(df["value"] - true_hv) / true_hv
ax = sns.lineplot(x="samples", y="value", hue="Method", data=df, marker="o")
ax.set(xscale="log", yscale="log", ylabel="Relative error")
plt.show()
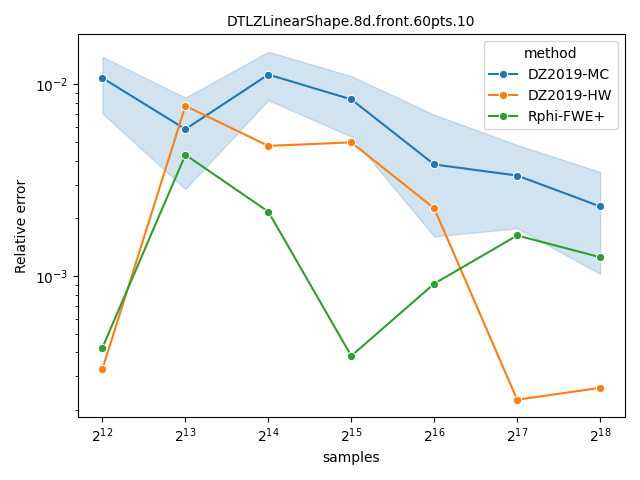
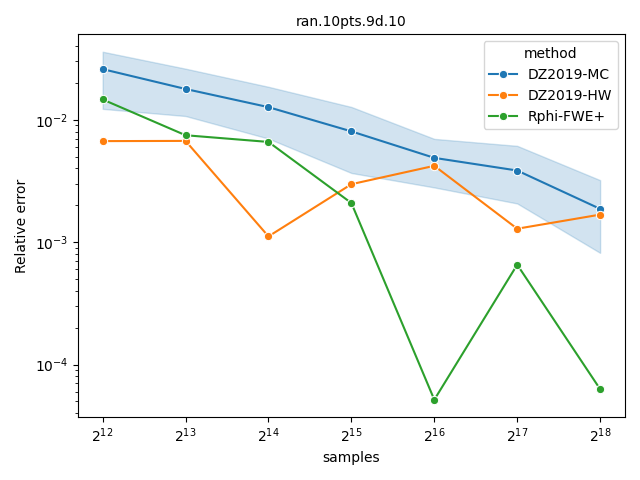
Comparing Monte-Carlo and quasi-Monte-Carlo approximations#
The quasi-Monte-Carlo approximations with method="Rphi-FWE+"
[14] and method="DZ2019-HW"
[13] is deterministic, but not monotonic on the
number of samples. Nevertheless, they are often better than the Monte-Carlo
approximation produced by method="DZ2019-MC"
[13], specially with large number of objectives. A
more detailed comparison is provided by López-Ibáñez[14].
datasets = ["DTLZLinearShape.8d.front.60pts.10", "ran.10pts.9d.10"]
ref = 10
samples = 2 ** np.arange(12, 19)
maxiter = samples.max()
for dataset in datasets:
x = moocore.get_dataset(dataset)[:, :-1]
x = moocore.filter_dominated(x)
exact = moocore.hypervolume(x, ref=ref)
res = []
for i in samples:
hv = moocore.hv_approx(x, ref=ref, nsamples=i, method="DZ2019-HW")
res.append(dict(samples=i, method="DZ2019-HW", hv=hv))
hv = moocore.hv_approx(x, ref=ref, nsamples=i, method="Rphi-FWE+")
res.append(dict(samples=i, method="Rphi-FWE+", hv=hv))
for k in range(5):
hv = moocore.hv_approx(
x, ref=ref, nsamples=i, method="DZ2019-MC", seed=k
)
res.append(dict(samples=i, method="DZ2019-MC", hv=hv))
df = pd.DataFrame(res)
df["hverror"] = np.abs(1.0 - (df.hv / exact))
df["method"] = (
df["method"]
.astype("category")
.cat.reorder_categories(["DZ2019-MC", "DZ2019-HW", "Rphi-FWE+"])
)
plt.figure()
ax = sns.lineplot(
data=df, x="samples", y="hverror", hue="method", marker="o"
)
ax.set_title(f"{dataset}", fontsize=10)
ax.set(yscale="log", ylabel="Relative error")
ax.set_xscale("log", base=2)
plt.tight_layout()
plt.show()
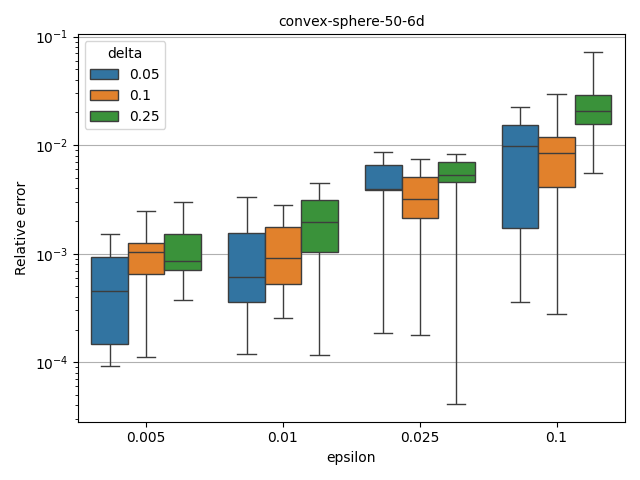
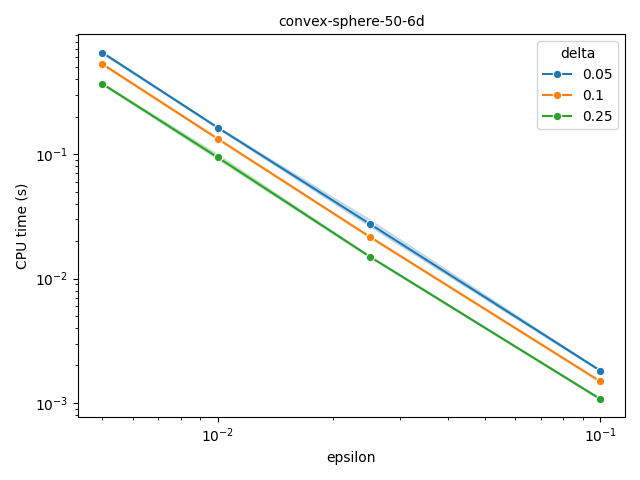
Fully polynomial-time randomized approximation scheme (FPRAS)#
moocore.hv_approx_fpras() allows obtaining an approximation with
relative error smaller than \(\epsilon\) (epsilon) with a given
probability \(1-\delta\) (delta). As the plot below shows, the
approximation error is often better than the requested value, but the
computation time increases very quickly for smaller epsilon.
shape = "convex-sphere"
ref = 1.1
npoints = 50
dim = 6
reps = 10
rng = np.random.default_rng(42)
res = []
for r in range(reps):
z = moocore.generate_ndset(npoints, dim, method=shape, seed=42 + r)
t_start = time.perf_counter()
exact = moocore.hypervolume(z, ref=ref)
t_end = time.perf_counter() - t_start
for epsilon in [0.1, 0.025, 0.01, 0.005]:
for delta in [0.25, 0.1, 0.05]:
t_start = time.perf_counter()
hv = moocore.hv_approx_fpras(
z, ref=ref, seed=rng, epsilon=epsilon, delta=delta
)
t_end = time.perf_counter() - t_start
res.append(
dict(
r=r,
epsilon=epsilon,
delta=delta,
hverror=np.abs(1 - hv / exact),
time=t_end,
)
)
df = pd.DataFrame(res)
df["delta"] = df["delta"].astype("category")
df["epsilon"] = df["epsilon"].astype("category")
plt.figure()
ax = sns.boxplot(df, x="epsilon", y="hverror", hue="delta", whis=[0, 100])
ax.set_title(f"{shape}-{npoints}-{dim}d", fontsize=10)
ax.set(yscale="log", ylabel="Relative error")
ax.yaxis.grid(True)
plt.tight_layout()
plt.figure()
ax = sns.lineplot(data=df, x="epsilon", y="time", hue="delta", marker="o")
ax.set_title(f"{shape}-{npoints}-{dim}d", fontsize=10)
ax.set(yscale="log", ylabel="CPU time (s)")
ax.set_xscale("log", base=10)
plt.tight_layout()
plt.show()
Total running time of the script: (0 minutes 38.394 seconds)