The following plots compare the performance of moocore
against emoa and
bbotk.
Other R packages are not included in the comparison because they are
based on these packages for the functionality benchmarked, so they are
at least as slow as them. For example GPareto,
mlr3mbo,
rmoo
and bbotk
use emoa to
compute the hypervolume. Not all packages provide the same
functionality.
Show benchmarking setup code
library(matrixStats)
library(data.table)
#>
#> Attaching package: 'data.table'
#> The following object is masked from 'package:base':
#>
#> %notin%
library(ggplot2)
library(moocore)
geomspace <- function(start, stop, num) {
x <- round(10^(seq(log10(start), log10(stop), length.out = num)), 0)
x[1] <- start
x[length(x)] <- stop
x
}
read_ndset <- function(filename, filter=FALSE) {
cat("Get file '", filename, "'\n")
destfile <- system.file(file.path("extdata", filename), package="moocore")
if (destfile == "") {
destfile <- file.path("../../../testsuite/data", filename)
if (!file.exists(destfile)) {
fileext <- if (endsWith(destfile, ".xz")) ".xz" else ""
destfile <- withr::local_tempfile(fileext = fileext)
base_url <- "https://github.com/multi-objective/testsuite/raw/refs/heads/main/data/"
utils::download.file(paste0(base_url, filename), destfile, quiet = FALSE)
}
}
x <- read_datasets(destfile)
x <- x[, -ncol(x)] # Union of datasets
if (filter)
x <- filter_dominated(x)
x
}
# This is adapted from atime:::plot.atime
benchmark_plot <- function (x, title = "", only_seconds=TRUE, ...) {
expr.name <- N <- kilobytes <- NULL
meas <- x[["measurements"]]
by.dt <- meas[, x$by.vec, with = FALSE]
tall.list <- list()
for (unit.i in seq_along(x$unit.col.vec)) {
col.name <- x$unit.col.vec[[unit.i]]
unit <- names(x$unit.col.vec)[[unit.i]]
if (is.null(unit) || unit == "")
unit <- col.name
tall.list[[unit.i]] <- meas[, data.table(N, by.dt,
unit, median = get(col.name))]
}
tall <- rbindlist(tall.list)
if (only_seconds) {
tall <- tall[unit=="seconds", ]
ylab <- "CPU time (seconds)"
} else {
ylab <- "median line, min/max band"
}
gg <- ggplot() + theme_bw(base_size=12) +
geom_ribbon(aes(N, ymin = min, ymax = max, fill = expr.name),
data = data.table(meas, unit = "seconds"), alpha = 0.25, show.legend=FALSE) +
geom_line(aes(N, median, color = expr.name), data = tall) +
geom_point(aes(N, median, color = expr.name), data = tall) +
scale_x_log10(breaks = unique(tall$N), labels = unique(tall$N)) +
scale_y_log10(ylab, labels = scales::label_log(), guide="axis_logticks") +
labs(subtitle = title) +
theme(axis.text.x = element_text(angle = 45, vjust = 1, hjust = 1))
if (only_seconds) {
gg <- gg + theme(legend.title = element_blank(),
legend.background = element_rect(fill=alpha("white", alpha=0.5)),
legend.key = element_rect(fill = NA, color = NA),
legend.position = "inside",
legend.position.inside = c(0.99, 0.01),
legend.justification = c("right", "bottom"),
legend.margin = margin_auto(0))
} else {
gg <- gg + theme(legend.title = element_blank(),
legend.background = element_rect(fill=alpha("white", alpha=0.5)),
legend.position = c(0.8, 0.625)) + facet_grid(unit ~ ., scales = "free")
}
gg
}
get_package_version <- function(package)
paste0(package, " (", as.character(packageVersion(package)), ")")
benchmark <- function(name, x, N, setup, expr.list, prefix, title) {
rds_file <- paste0("bench/bench-", prefix, "-", name, ".rds")
if (run_benchmarks || !file.exists(rds_file)) {
lapply(names(expr.list), library, character.only = TRUE)
names(expr.list) <- sapply(names(expr.list), get_package_version, USE.NAMES=FALSE)
res <- substitute(atime::atime(
N = N,
expr.list = expr.list,
setup = SETUP,
result=FALSE,
times=5,
seconds.limit=10), list(SETUP=setup))
res <- eval(res)
saveRDS(res, file = rds_file)
} else {
res <- readRDS(rds_file)
}
gg <- benchmark_plot(res, title = paste0(title, " for ", name))
gg
}
get_ndset <- function(x, filter)
{
if (is.null(x$generate)) return(read_ndset(x$file, filter=filter))
do.call(moocore::generate_ndset, x$generate)
}
benchmark_all <- function(files, prefix, title, setup, expr.list, filter)
{
for (name in names(files)) {
p <- benchmark(name = name, x = get_ndset(files[[name]], filter=filter),
N = files[[name]]$N, prefix = prefix, title = title,
setup = setup, expr.list = expr.list)
print(p)
}
}Identifying (non)dominated points
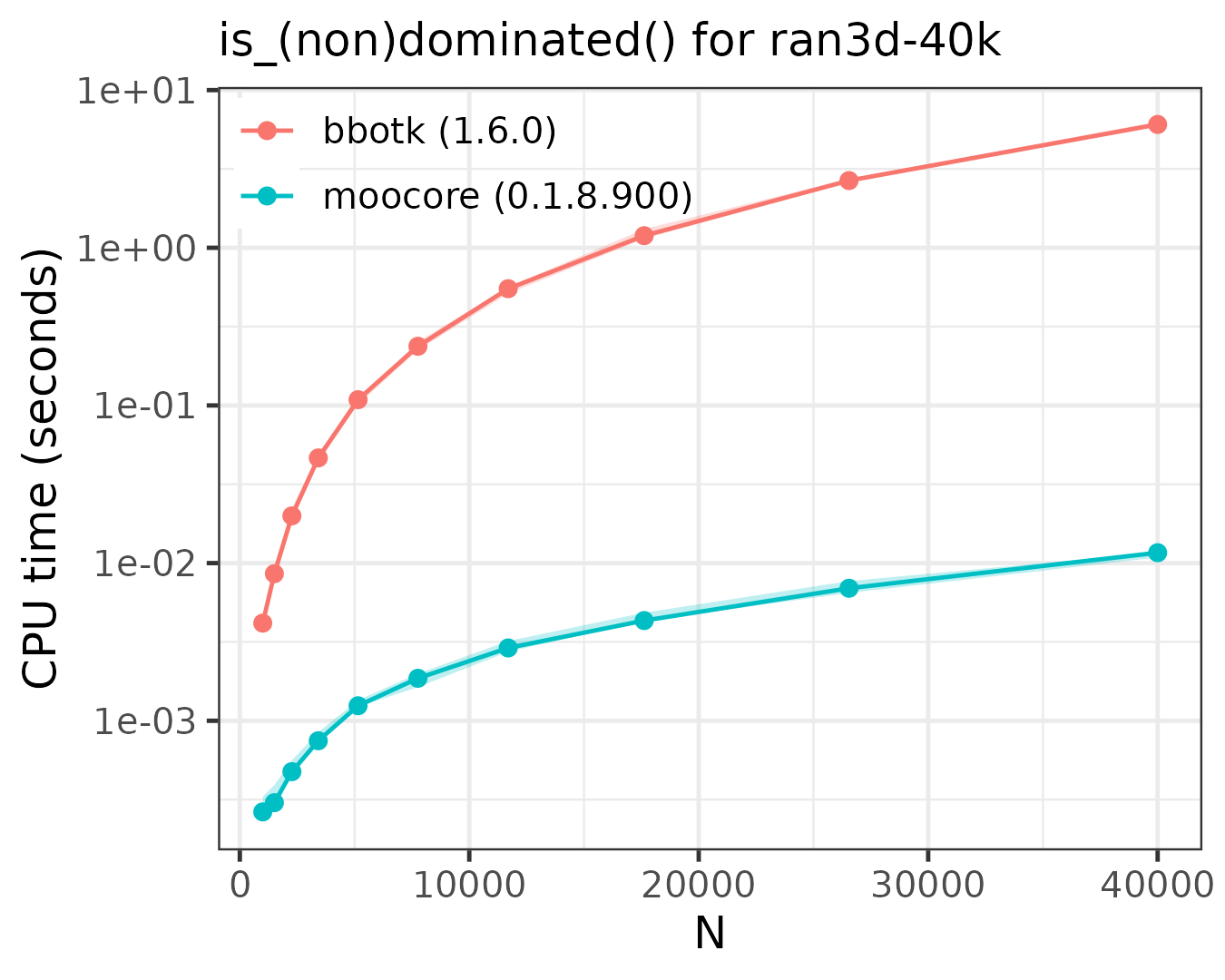
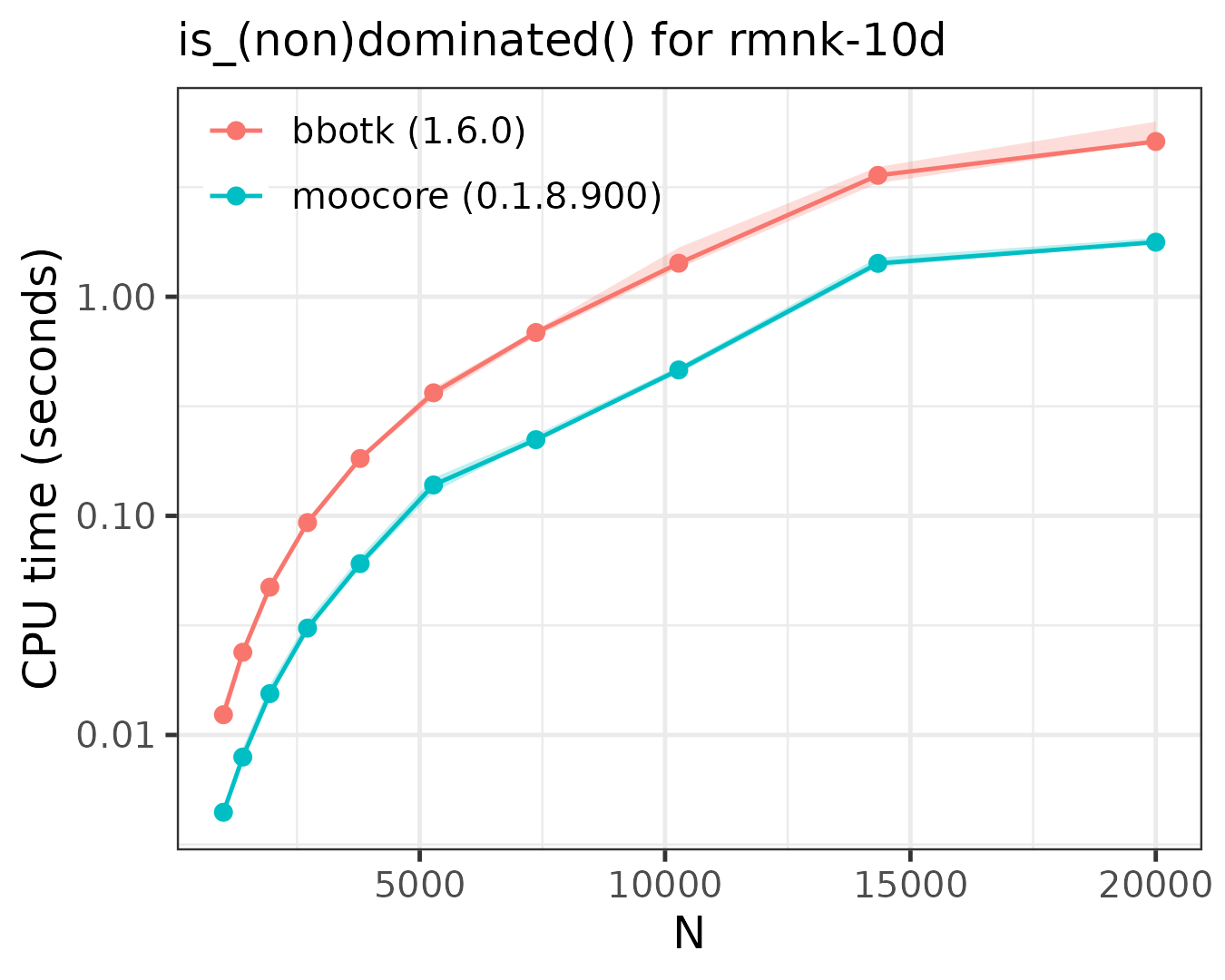
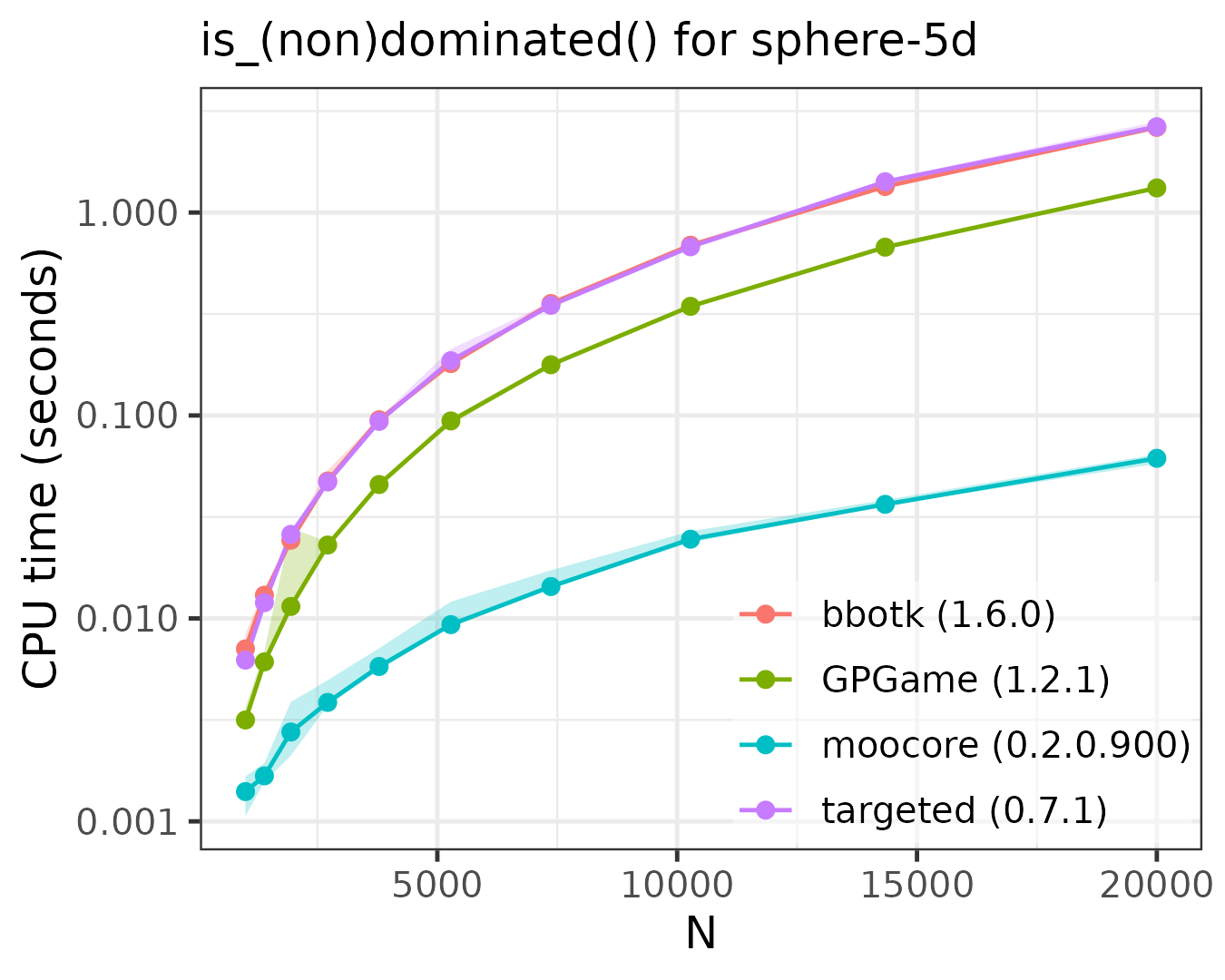
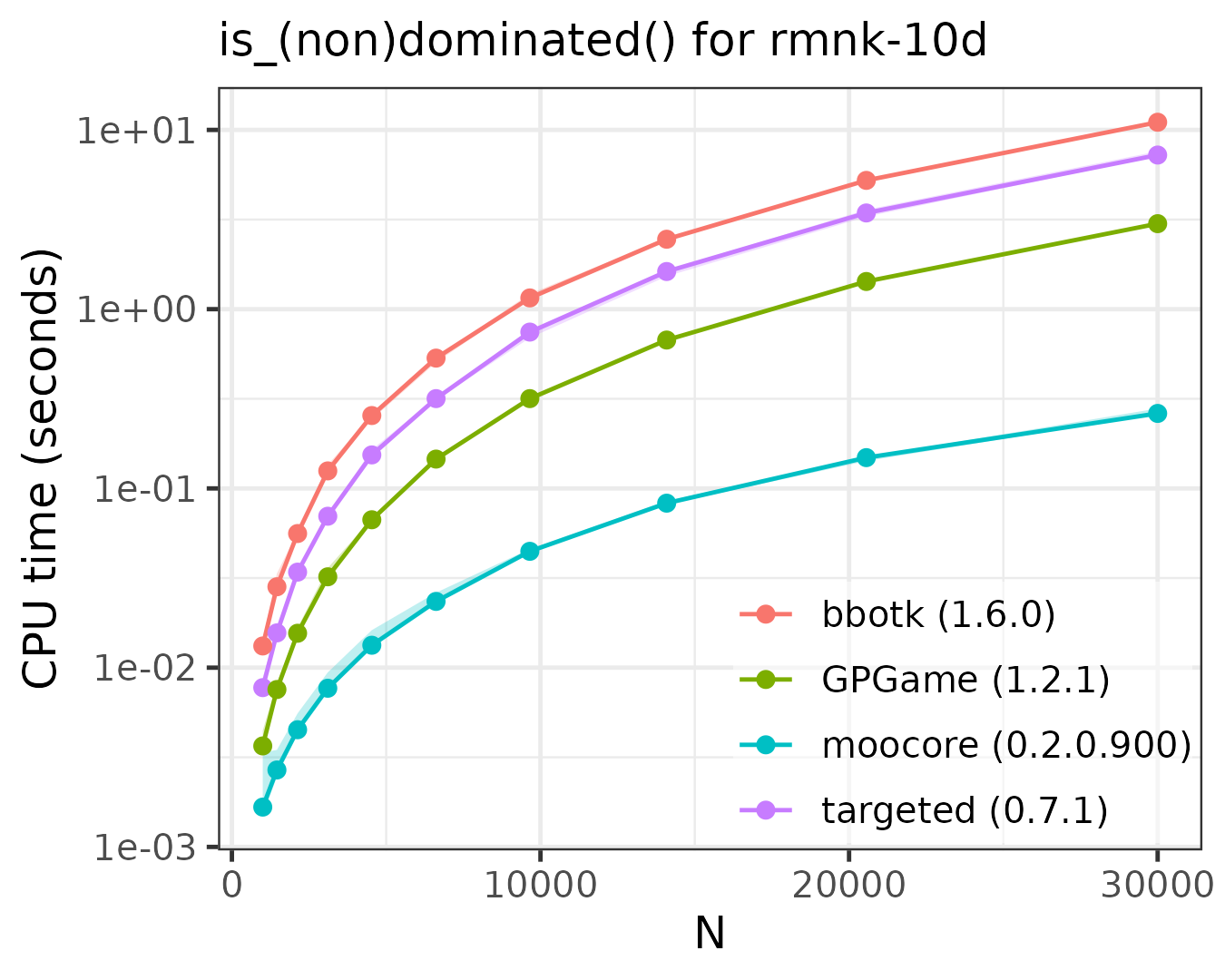
The following plots compare the speed of finding (non)dominated
solutions, equivalent to moocore::is_nondominated(), in 2D,
3D, 4D, 5D and 10D. The plots show that moocore
is always 10 times faster than GPGame,
targeted
and bbotk
for any number of objectives. In addition, GPGame
calculates wrong values with repeated coordinates (vpicheny/GPGame#2).
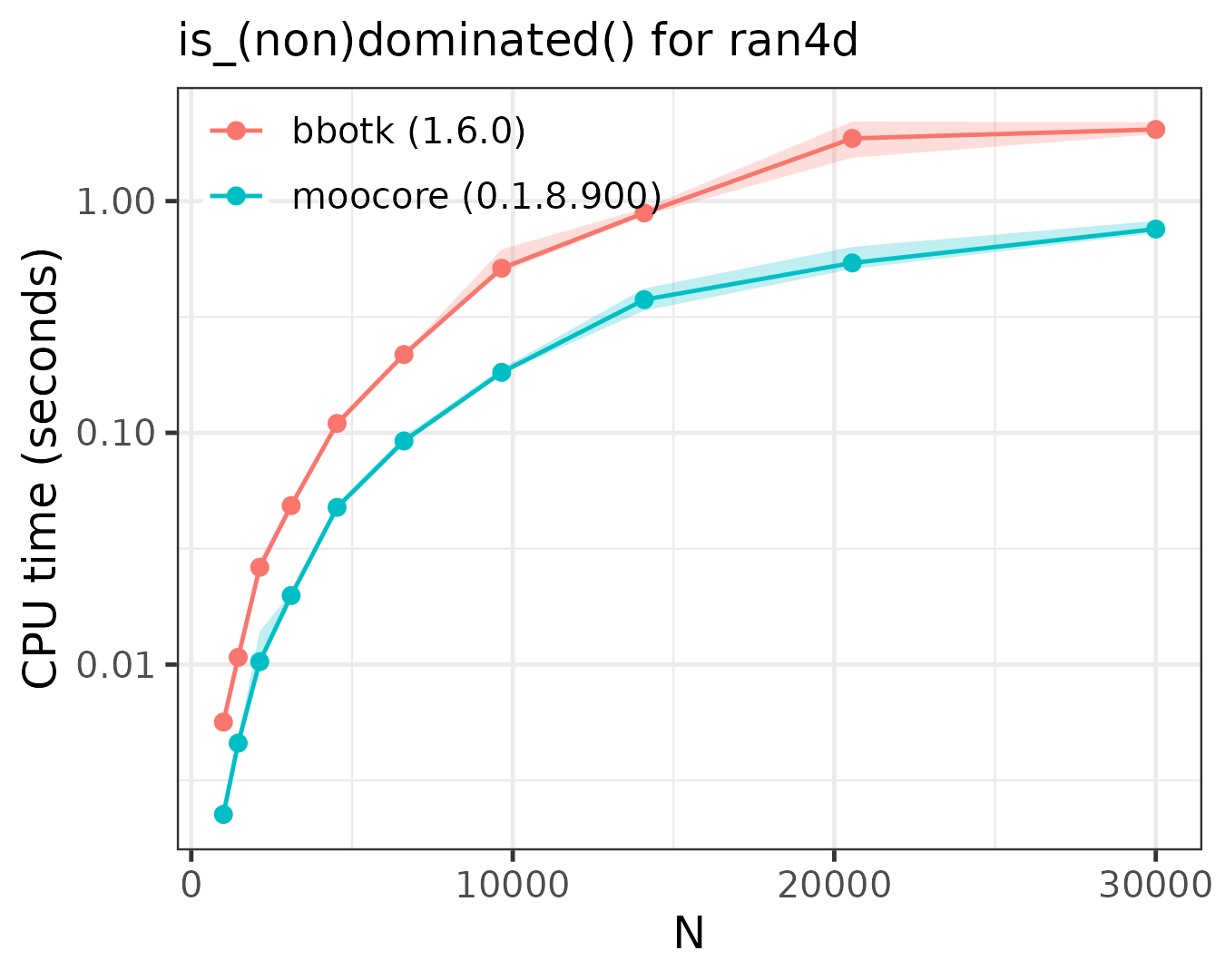
The rand4d testcase is somewhat different from the others,
as it is mostly composed of dominated points (only 10590 points are
nondominated), while the other testcases are mostly nondominated
sets.
setup <- quote({
stopifnot(nrow(x) >= N)
z <- as.matrix(x[1L:N, ])
tz <- t(z)
})
expr.list <- list(
moocore = quote(which(moocore::is_nondominated(z))),
bbotk = quote(which(bbotk::is_dominated(tz))),
GPGame = quote(GPGame::nonDom(z)),
targeted = quote(targeted::nondom(z))
)
files <- list(
"test2D-200k"=list(file="test2D-200k.inp.xz", N=geomspace(1000, 50000, 10)),
"ran3d-40k"=list(file="ran.40000pts.3d.1.xz", N=geomspace(1000, 25000, 10)),
"ran4d"=list(file="ran.9000pts.4d.10.xz", N=geomspace(1000, 60000, 10)),
"convex-4d"=list(generate=list(25000, 4, "convex-sphere", 42), N=geomspace(500, 25000, 10)),
"sphere-5d"=list(generate=list(20000, 5, "sphere", 42), N=geomspace(100, 20000, 10)),
"rmnk-10d"=list(file="rmnk_0.0_10_16_1_0_ref.txt.xz", N=geomspace(100, 20000, 10))
)
benchmark_all(files, prefix="ndom", title = "is_(non)dominated()",
setup = setup, expr.list = expr.list, filter=FALSE)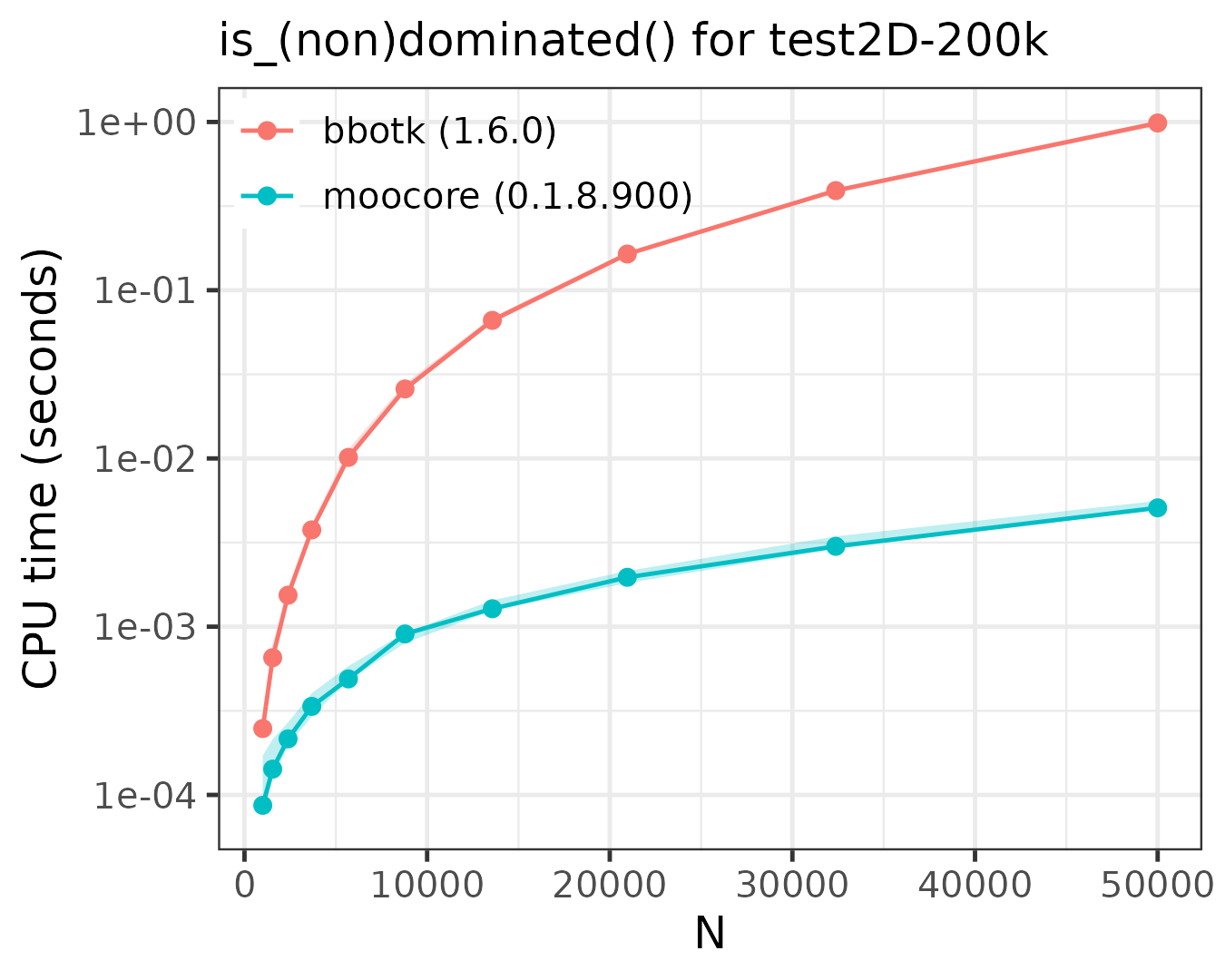
Exact computation of hypervolume
The following plots compare the speed of computing the hypervolume indicator in 3D, 4D, 5D and 6D.
setup <- quote({
ref <- colMaxs(x, useNames = FALSE) + 1L
stopifnot(nrow(x) >= N)
z <- x[1L:N, ]
tz <- t(z)
})
expr.list <- list(
moocore = quote(moocore::hypervolume(z, ref = ref)),
emoa = quote(emoa::dominated_hypervolume(tz, ref = ref)))
files <- list(
"DTLZLinearShape.3d"=list(
file = "DTLZSphereShape.3d.front.1000pts.10",
N = geomspace(200, 6500, 10)),
"DTLZLinearShape.4d"=list(
file = "DTLZLinearShape.4d.front.1000pts.10",
N = geomspace(100, 2000, 10)),
"DTLZLinearShape.5d"=list(
file = "DTLZLinearShape.5d.front.500pts.10",
N = geomspace(100, 1000, 10)),
"DTLZLinearShape.6d"=list(
file = "DTLZLinearShape.6d.front.700pts.10.xz",
N = geomspace(100, 1000, 10))
)
benchmark_all(files, prefix="hv", title = "HV Computation",
setup = setup, expr.list = expr.list, filter=TRUE)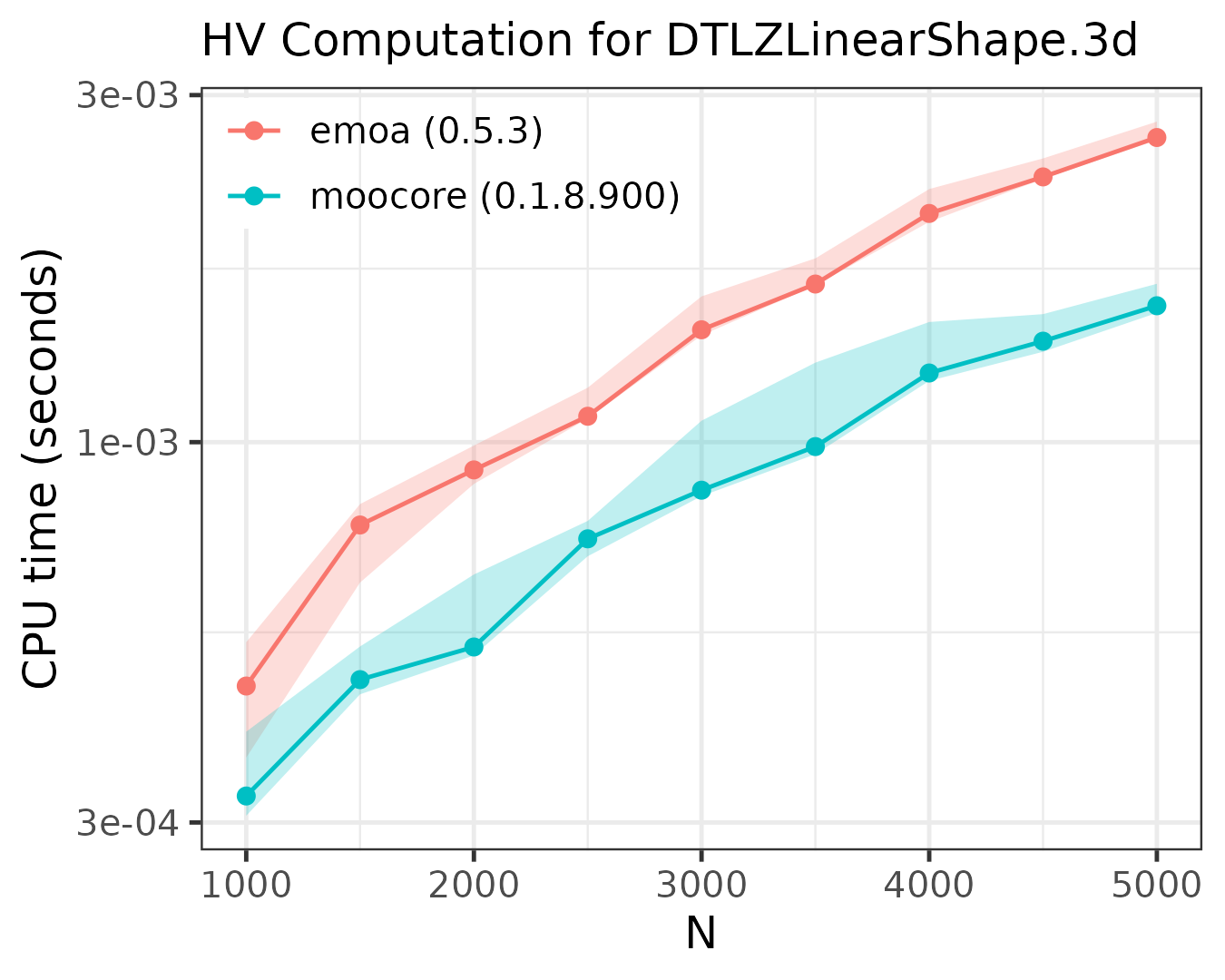
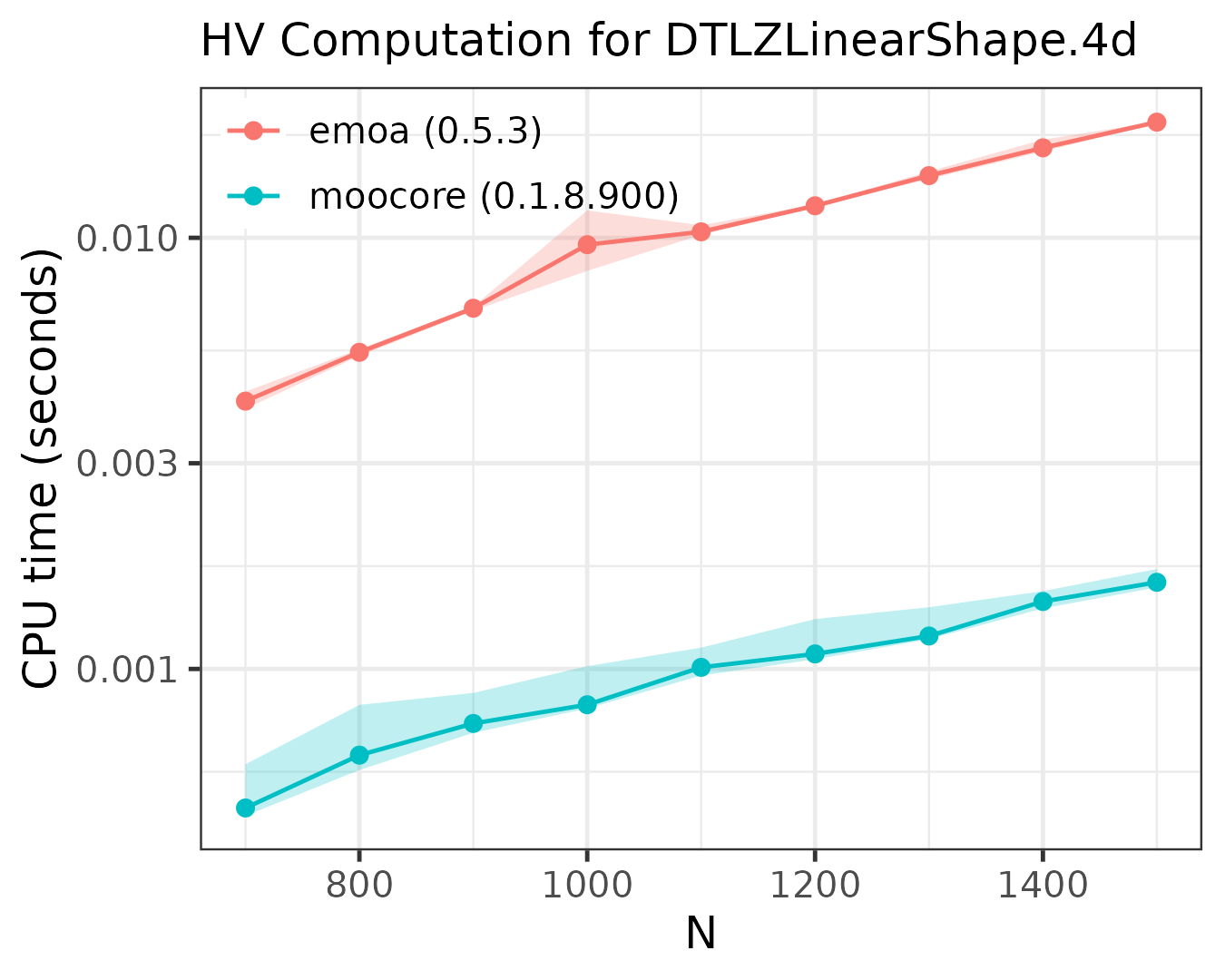
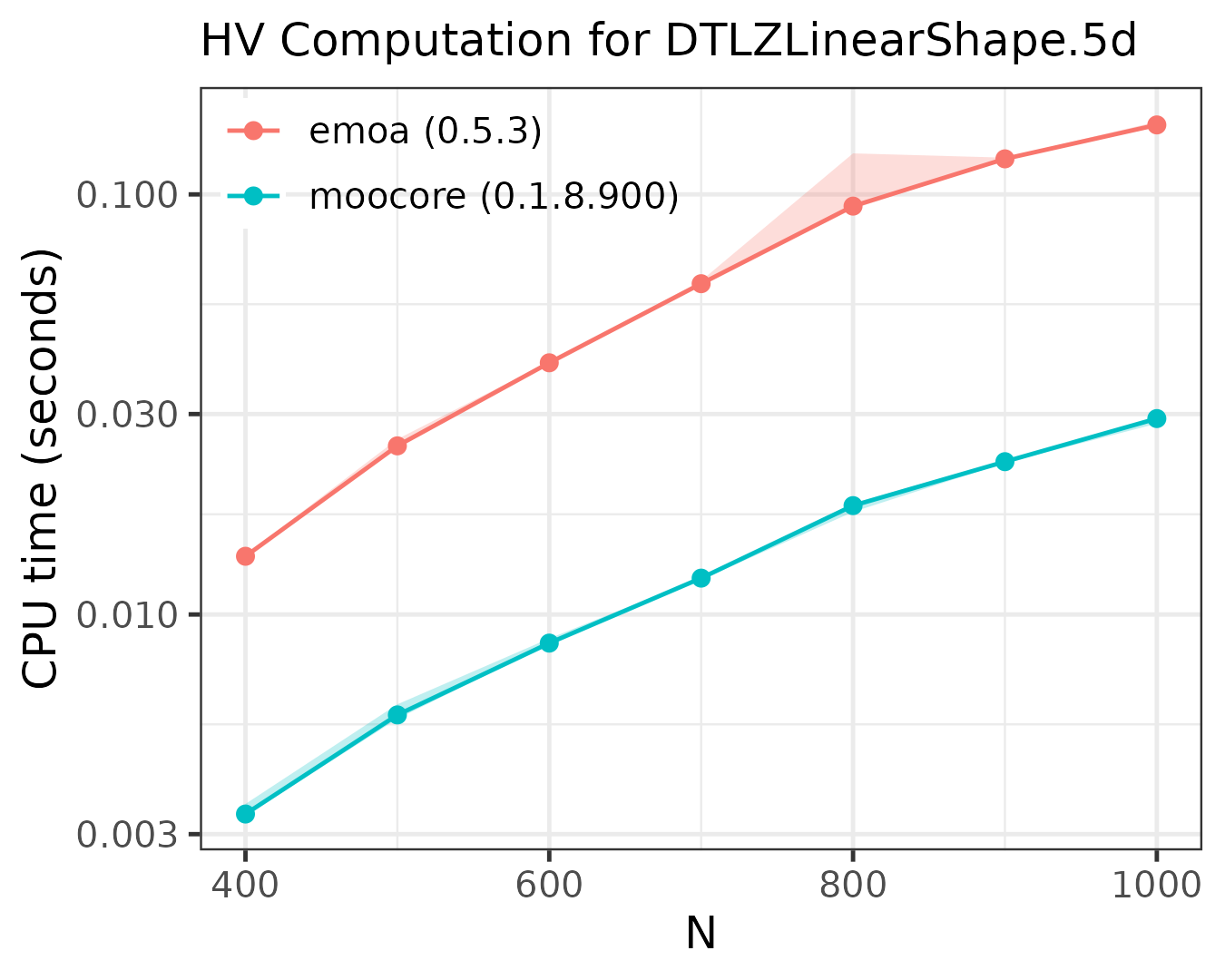
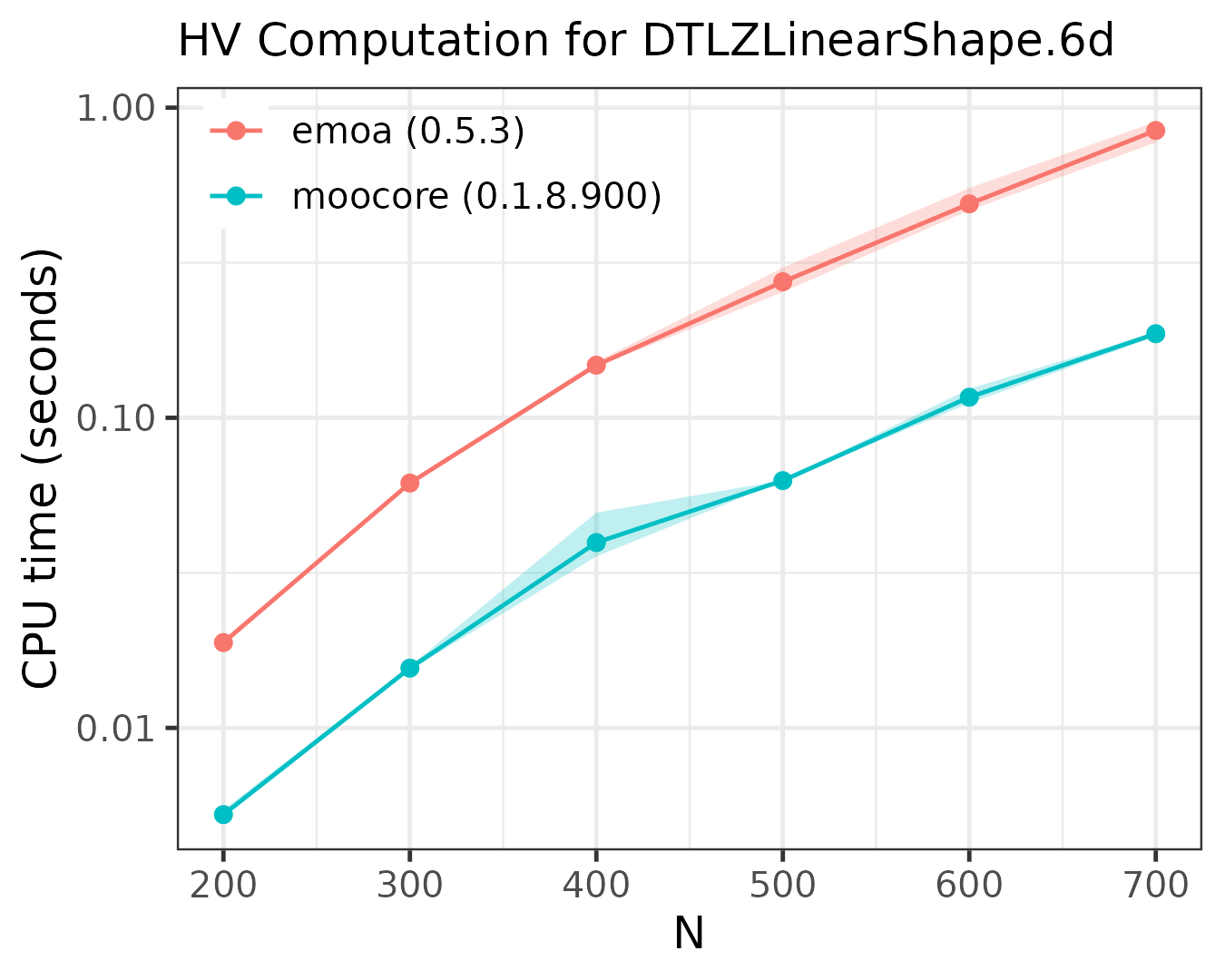
As the plots show, moocore
is always faster than emoa
and, hence, faster than GPareto,
mlr3mbo,
rmoo
and bbotk.
Hypervolume contribution
The only R package, other than moocore,
able to compute hypervolume contributions
moocore::hv_contributions() is emoa.
However, emoa is
buggy and calculates wrong values (olafmersmann/emoa#1).