Rank points according to Pareto-optimality (nondominated sorting).
Source:R/nondominated.R
pareto_rank.Rdpareto_rank() is meant to be used like rank(), but it assigns ranks
according to Pareto dominance, where rank 1 indicates those solutions not
dominated by any other solution in the input set. Duplicated points are
assigned the same rank. The resulting ranking can be used to partition
points into a list of matrices, each matrix representing a nondominated
front (Deb et al. 2002)
(see examples below).
Arguments
- x
matrix()|data.frame()
Matrix or data frame of numerical values, where each row gives the coordinates of a point.- maximise
logical()
Whether the objectives must be maximised instead of minimised. Either a single logical value that applies to all objectives or a vector of logical values, with one value per objective.
Value
An integer vector of the same length as the number of rows of the
input x, where each value gives the rank of each point (lower is
better).
Details
Given a finite set of points \(X \subset \mathbb{R}^m\), the rank of a point \(x \in X\) is defined as:
$$\operatorname{rank}(x) = r \iff x \in F^c_{r} \land \nexists y \in F^c_{r}, y \prec x$$
where \(y \prec x\) means that \(y\) dominates \(x\) according to Pareto optimality, \(F^c_r = X \setminus \bigcup_{i=1}^{r-1} F_i\) and \(F_r = \{x \in X \land \operatorname{rank}(x) = r\}\). The sets \(F_c\), with \(c=1,\dots,k\), partition \(X\) into \(k\) fronts, that is, mutually nondominated subsets of \(X\).
With \(m=2\), i.e., ncol(data)=2, the code uses the best-known
\(O(n \log n)\) algorithm by Jensen (2003)
. When \(m \geq 3\), it
uses the naive algorithm that identifies one front at a time, which
requires \(O(n^2\log n)\) for \(m=3\), and \(O(n^2 \log^{m-2} n)\)
for \(m \geq 4\).
References
Kalyanmoy Deb, A Pratap, S Agarwal, T Meyarivan (2002).
“A fast and elitist multi-objective genetic algorithm: NSGA-II.”
IEEE Transactions on Evolutionary Computation, 6(2), 182–197.
doi:10.1109/4235.996017
.
M
T Jensen (2003).
“Reducing the run-time complexity of multiobjective EAs: The NSGA-II and other algorithms.”
IEEE Transactions on Evolutionary Computation, 7(5), 503–515.
Examples
three_fronts = matrix(c(1, 2, 3,
3, 1, 2,
2, 3, 1,
10, 20, 30,
30, 10, 20,
20, 30, 10,
100, 200, 300,
300, 100, 200,
200, 300, 100), ncol=3, byrow=TRUE)
pareto_rank(three_fronts)
#> [1] 1 1 1 2 2 2 3 3 3
split.data.frame(three_fronts, pareto_rank(three_fronts))
#> $`1`
#> [,1] [,2] [,3]
#> [1,] 1 2 3
#> [2,] 3 1 2
#> [3,] 2 3 1
#>
#> $`2`
#> [,1] [,2] [,3]
#> [1,] 10 20 30
#> [2,] 30 10 20
#> [3,] 20 30 10
#>
#> $`3`
#> [,1] [,2] [,3]
#> [1,] 100 200 300
#> [2,] 300 100 200
#> [3,] 200 300 100
#>
path_A1 <- file.path(system.file(package="moocore"),"extdata","ALG_1_dat.xz")
set <- read_datasets(path_A1)[,1:2]
ranks <- pareto_rank(set)
str(ranks)
#> int [1:23260] 13 24 20 22 22 5 23 5 16 20 ...
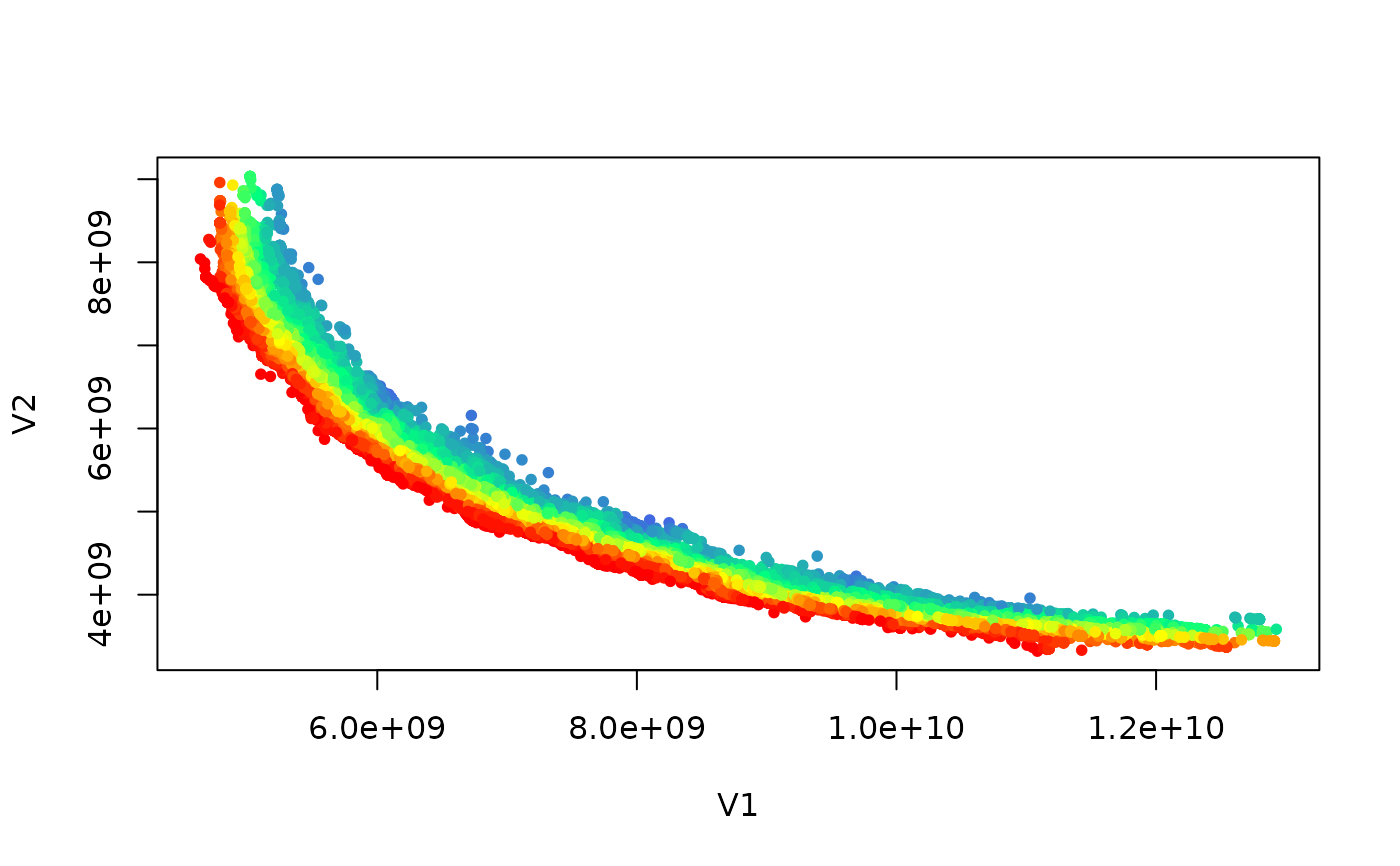
if (requireNamespace("graphics", quietly = TRUE)) {
colors <- colorRampPalette(c("red","yellow","springgreen","royalblue"))(max(ranks))
plot(set, col = colors[ranks], type = "p", pch = 20)
}